This kit applies ybbR peptide and Sfp enzyme to functionalize your protein of interest with DNA handles for subsequent optical-tweezers measurements of protein folding and conformational dynamics. The system constitutes an alternative to the Protein labeling and tethering kit (cysteine). The Protein tethering kit (ybbR) is the ideal option for applications where cysteine incorporation is not possible, for example, due to the presence of functional native cysteines in your protein of interest.
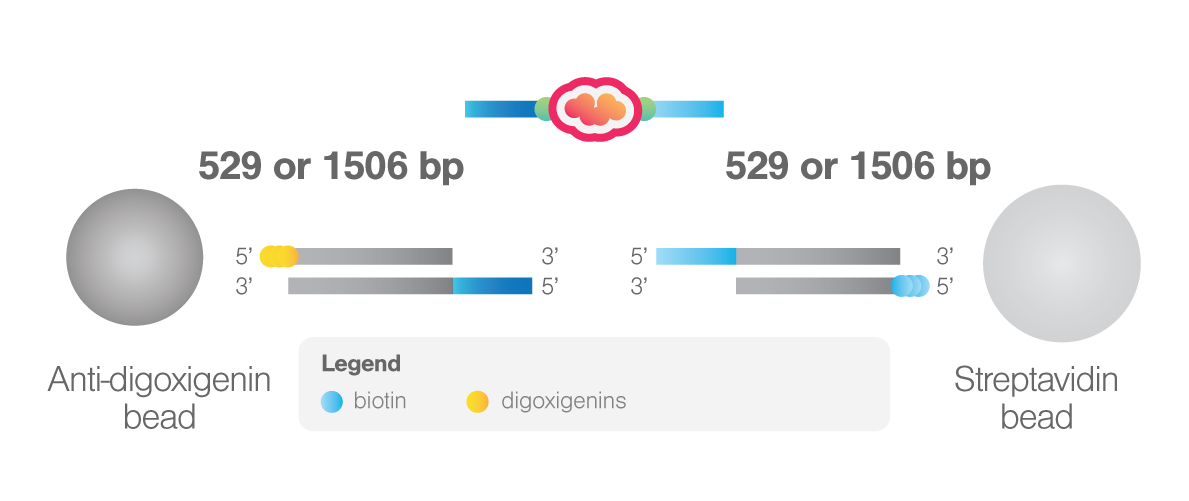
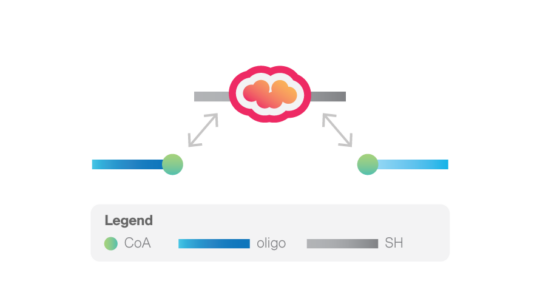
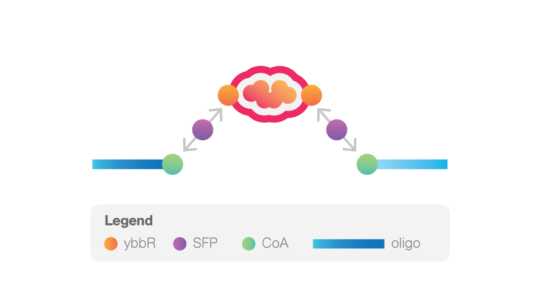
The system is based on the enzymatic reaction of Sfp synthease enzyme that catalyzes the covalent binding of a CoA-modified oligonucleotide to each of the two ybbR peptides inserted in the protein of interest. These oligonucleotides then anneal with the complementary 5’-overhangs of two DNA handles. For the DNA handles, two choices are available. The short handles (529 bp) will result in the highest force resolution, while the long handles (1506 bp) allow a more straightforward tethering procedure.
The kit also includes purified and functionalized adenosine kinase (AdK) fused to two ybbR peptides that can be used as a control protein for each step of the labeling procedure.
Our protein labeling and tethering kit (ybbR) relieves you from costs and time that are typically required for personnel training, troubleshooting, experiment optimization, and custom application development.
In case additional protein labelings with oligonucleotides or additional measurements with handles are required, CoA-oligonucleotides (including Sfp enzyme) or DNA handles can be ordered in separate packages.
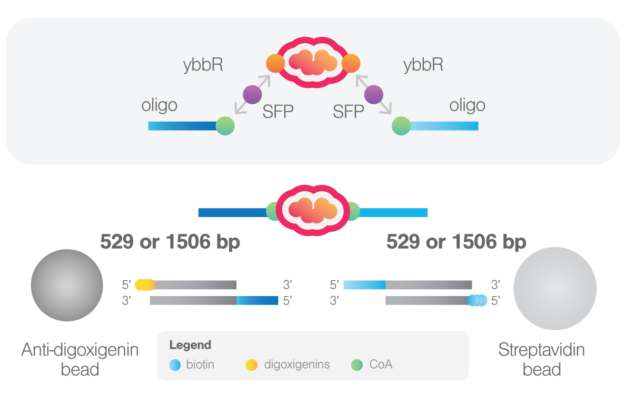
The kit enables 2 protein labelings with oligonucleotides (allowing up to 40 experimental sessions per labeled protein on the C-Trap) and supplies DNA handles for 10 experimental sessions on the C-Trap.
Estimated 1-4 weeks delivery time.