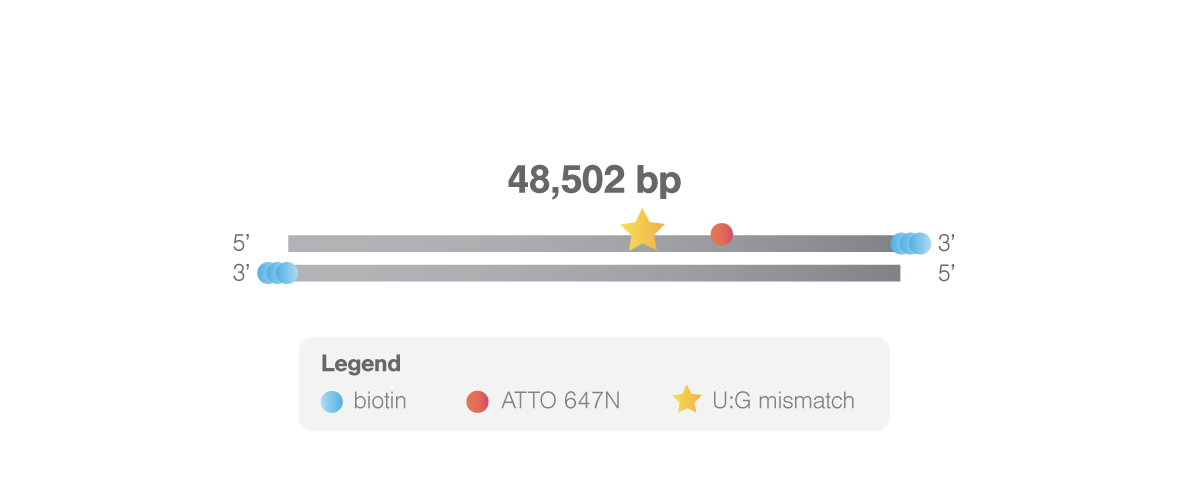
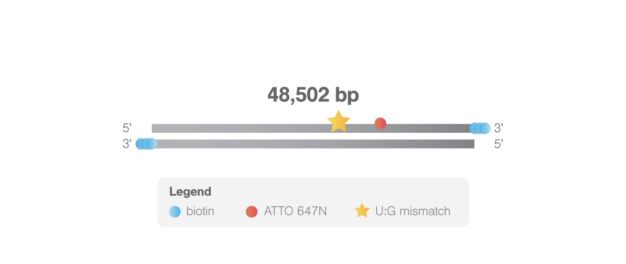
This biotinylated double-stranded λ-DNA (48,502 bp) is modified with a U:G mismatch at location 33,787 bp and an ATTO 647N fluorophore at location 44,826 bp. The lesion allows studying DNA mismatch repair mechanisms such as nucleotide excision repair and base excision repair. The fluorophore facilitates determination of the tether directionality, localization of the imaged protein in the DNA sequence, and can be used to bring the sample into focus. For visualizing the fluorophore, the 638/639 nm laser of the C-Trap is required.
The DNA is labeled with 3 biotins on each 3’ end to enable tethering in optical tweezers. The DNA lesion will be present in 90% of the tethers containing the fluorophore. Typically, 70% of the tethers contain both the lesion and the fluorophore.
The other available lesions are G:T mismatch, U:A mismatch and U:G mismatch. For request on other DNA lesions, please contact store@lumicks.com.
For the tethering procedure have a look at the protocol for tethering biotinylated λ-DNA (48,502 bp) and for the recommended buffers, have a look at the protocol for oxygen scavenging systems.
Estimated for 20 experimental sessions on the C-Trap.
Estimated 1-4 weeks delivery time.